Overcoming Replicative Limits: Mechanisms and Applications of Primary Cell Immortalization
Apr 21,2026
Primary cells retain key in vivo characteristics and are widely used in molecular, cellular, and biomedical research. However, their proliferative capacity is limited; after several passages, they enter senescence and eventually cease dividing, a phenomenon known as the Hayflick limit. Overcoming this constraint through immortalization is therefore a critical strategy.
In this issue of Cell Culture Academy, we review the molecular mechanisms of primary cell immortalization and outline core experimental approaches with representative examples.
I. Methods for Primary Cell Immortalization
Primary cell immortalization is commonly achieved by gene transfection, which introduces exogenous immortalizing genes into target cells to generate stable cell lines. These cells retain key primary cell characteristics while acquiring unlimited proliferative capacity. The most widely used approaches involve SV40 large T antigen (TAg) and telomerase reverse transcriptase (TERT).
1. SV40-Mediated Immortalization
Simian Virus 40 (SV40) encodes large T antigen, small T antigen, and structural proteins, with large T antigen serving as the principal driver of immortalization. It binds to p53 and related proteins, forming complexes that inactivate p53, disrupt tumor suppressor function, and prevent cell cycle arrest. This promotes cell cycle progression and enables cellular immortalization.
2. Telomerase-Mediated Immortalization
Telomeres are essential for chromosomal stability and progressively shorten with each cell division. Once they reach a critical length, cells enter senescence and cease proliferation. Telomerase, particularly its catalytic subunit TERT, maintains telomere length by reverse-transcribing RNA templates into telomeric DNA and adding it to chromosome ends. In most normal eukaryotic cells, telomerase activity is insufficient to sustain telomere length. Ectopic TERT expression restores telomerase activity, extends telomeres, and enables cellular immortalization.
Ⅱ. Example of Immortalized Primary Cells
Human umbilical vein endothelial cells (HUVECs) were used as target cells. SV40 TAg lentivirus was produced in 293T cells and used to infect HUVECs, followed by antibiotic selection to generate a stable SV40T-expressing immortalized cell. The workflow is outlined below:
1. Lentiviral Packaging
293T cells were seeded and transfected at 70-80% confluence with an SV40 TAg expression vector and lentiviral packaging plasmids.
After 48-72 h, the supernatant was collected.
The supernatant was centrifuged at 300-500 × g for 5 min at 4℃ and filtered through a 0.45 μm filter to remove debris. The viral supernatant was concentrated if necessary.
Aliquots were stored at -80℃.
2. Lentiviral Infection and Selection
HUVECs were seeded in 24-well plates at approximately 5 × 104 cells per well and cultured to 50-60% confluence.
The medium was replaced with fresh medium containing lentiviral particles.
Polybrene was added to a final concentration of 4-8 μg/mL to enhance infection efficiency.
After 6-8 h, the medium was replaced with fresh complete medium, and cells were cultured for an additional 24-48 h.
At 48 h post-infection, a selection antibiotic (e.g., G418, 1 mg/mL) was added. Cells were maintained under selection for 1-2 weeks, with medium changed every 2-3 days.
Non-transduced cells were eliminated, while transduced cells survived and formed stable resistant clones for expansion.
3. Cell Characterization
- Genomic Integration Analysis
Genomic DNA from immortalized HUVECs was analyzed by Sanger sequencing, confirming integration of the SV40 TAg gene into the host genome (Figure 1).
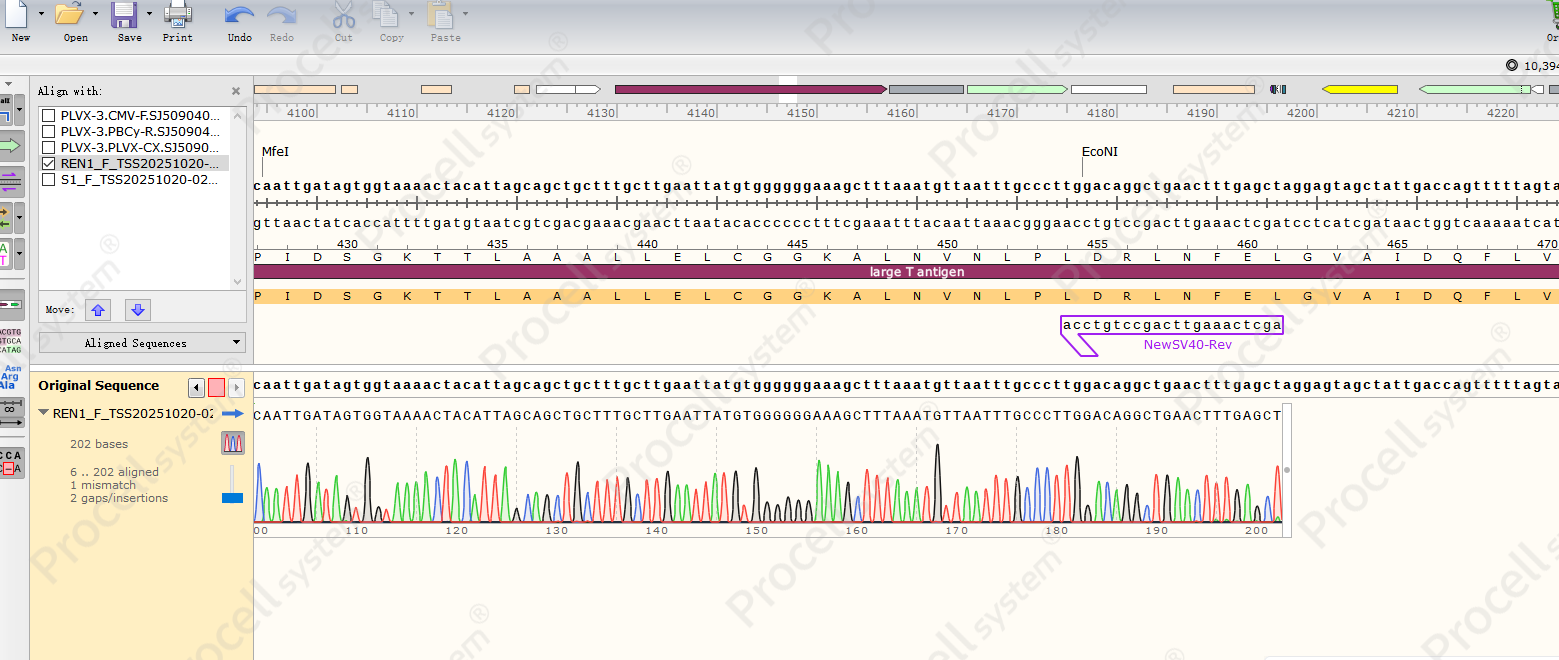
Figure 1. Sanger sequencing of SV40 TAg gene integration in immortalized HUVECs.
- Cell Proliferation Analysis
Immortalized HUVECs (HUVEC-SV40) showed increased proliferation compared with control cells (HUVEC-NC) (Figure 2) and maintained typical endothelial morphology through passage 25 (P25) (Figure 3).
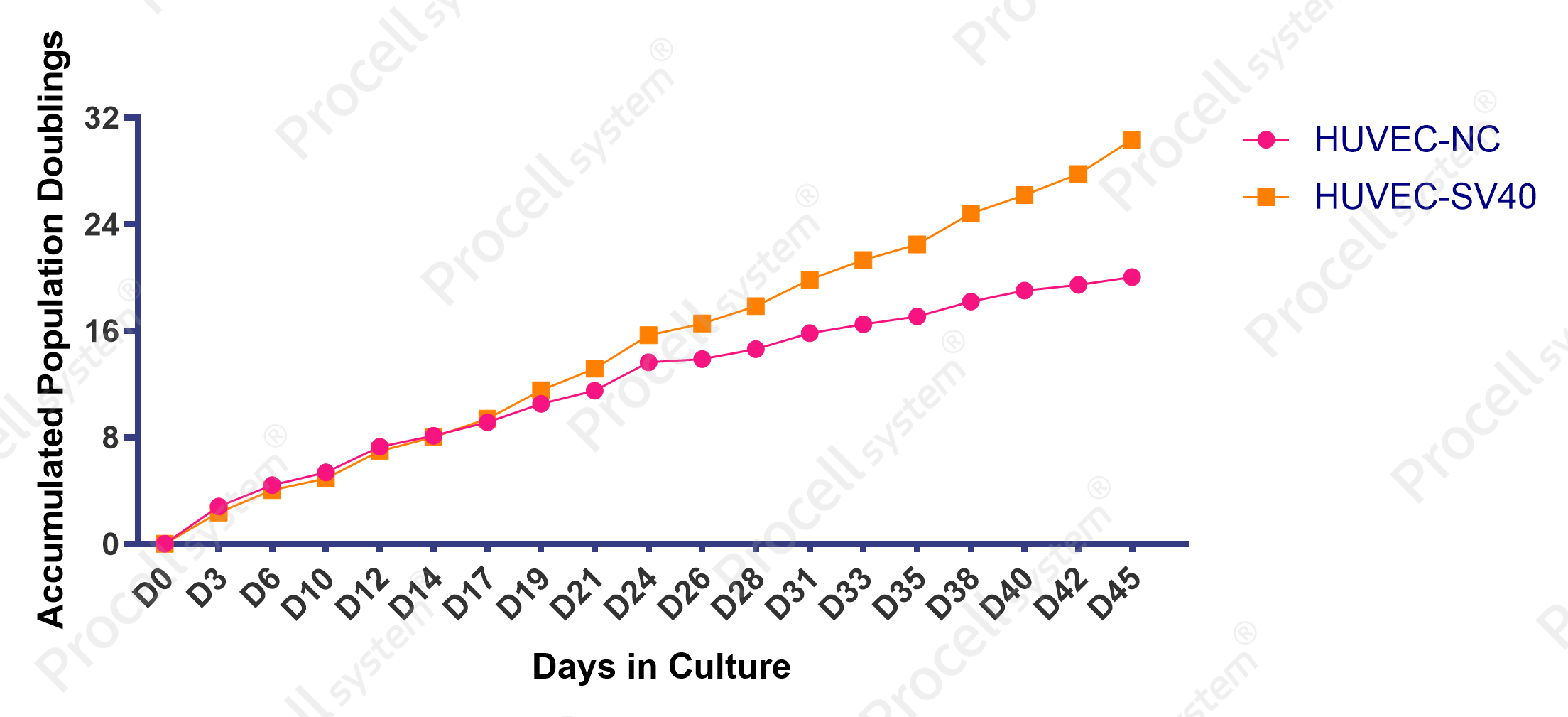
Figure 2. Proliferation of immortalized HUVECs
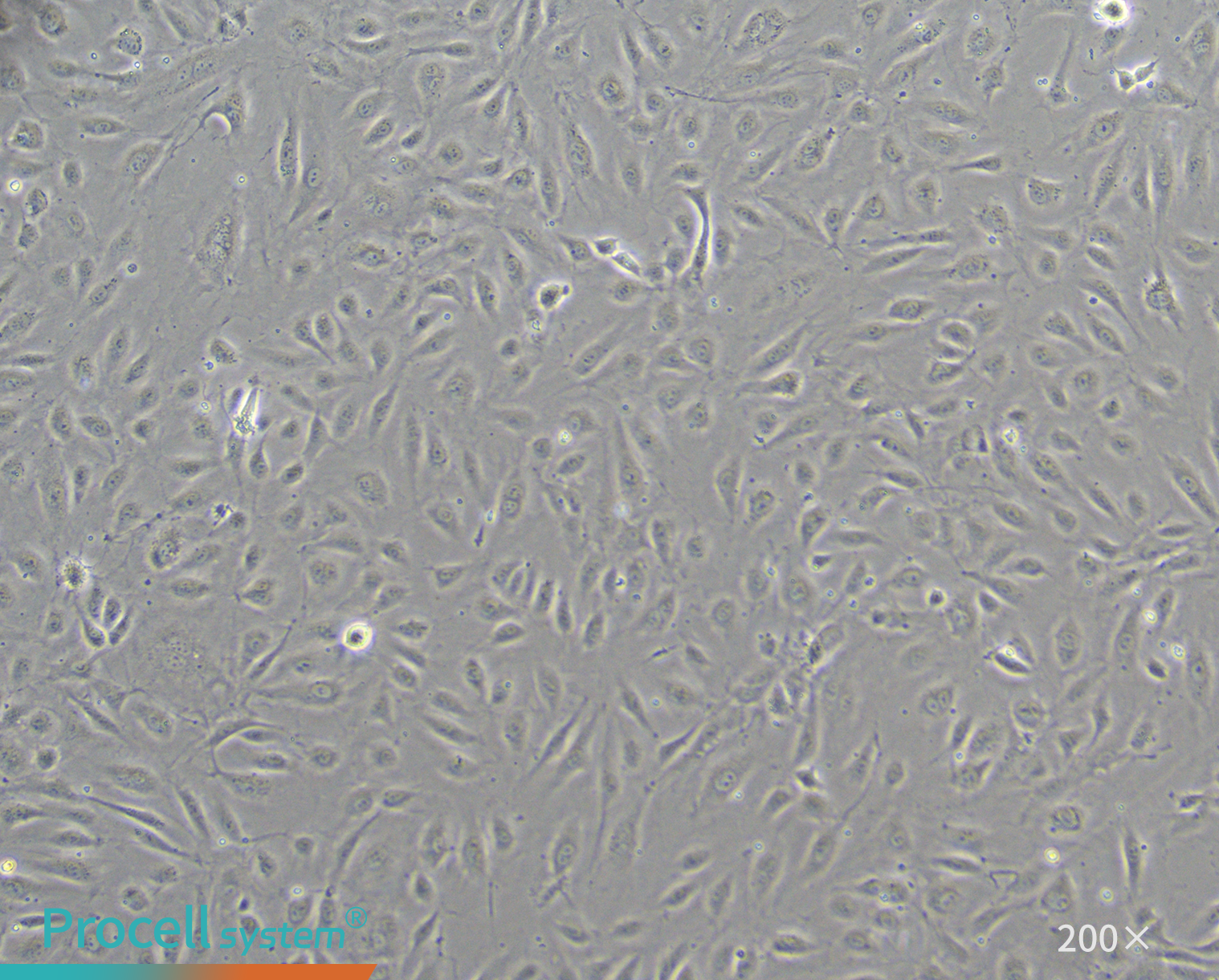
Figure 3. Morphology of immortalized HUVECs at P25
- Immunofluorescence Characterization
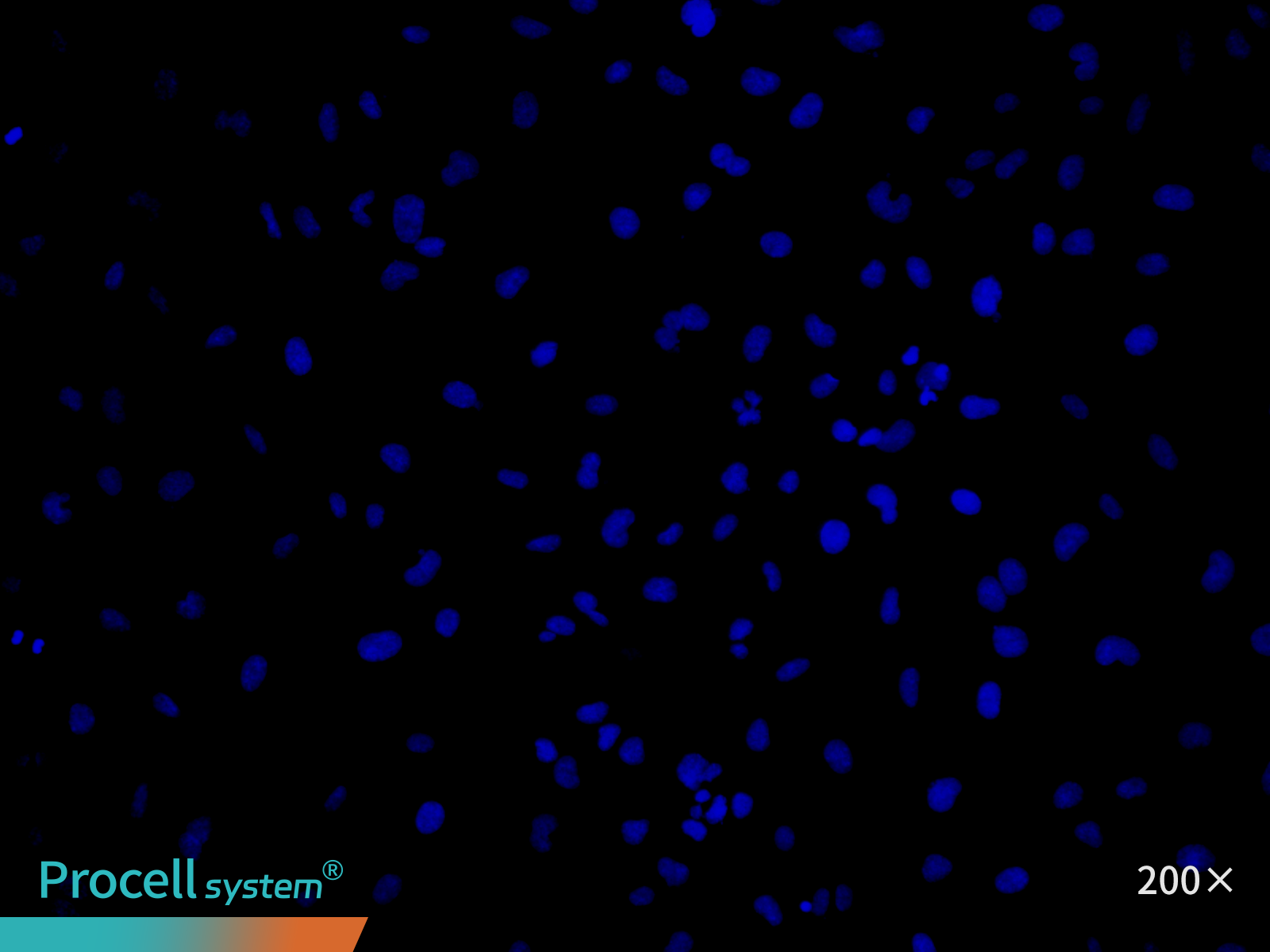
At P25, immunofluorescence staining for the endothelial marker CD31 indicated ≥ 90% cell-positive cells. (Figure 4).
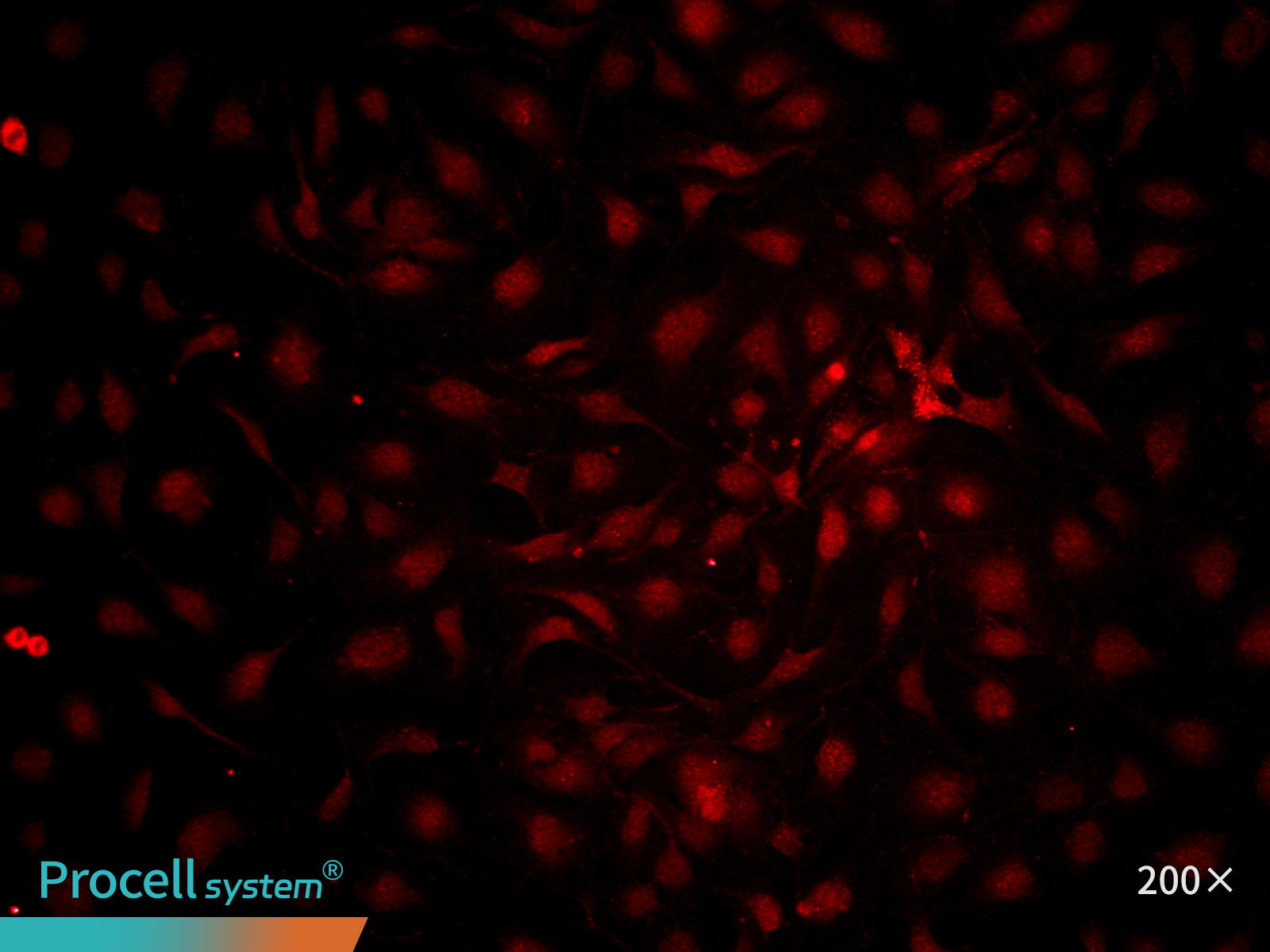 |
 |
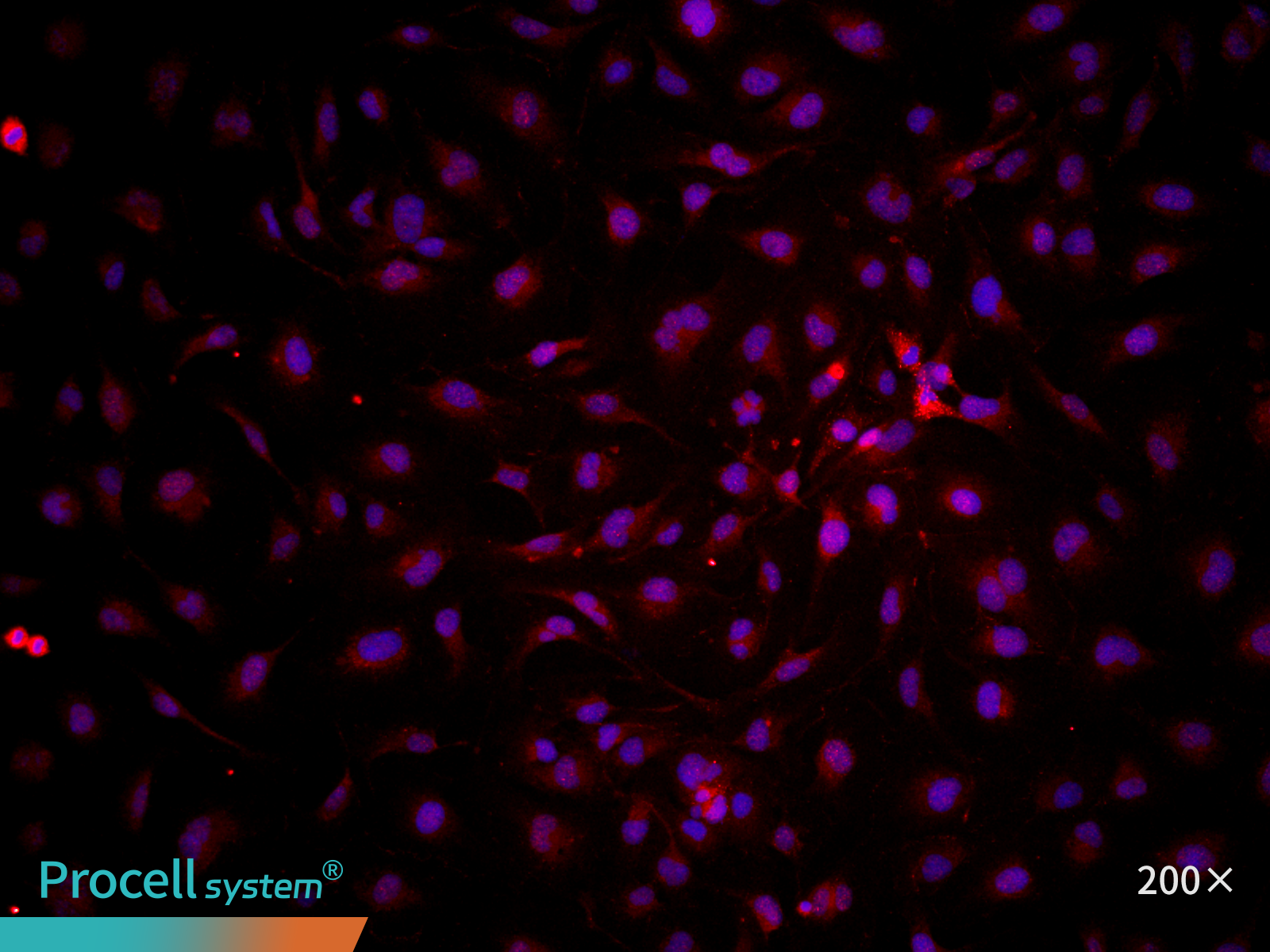 |
| CD31 | Hoechst | Merge |
Figure 4. Immunofluorescence analysis of immortalized HUVECs (P25)
- Functional Validation
Angiogenic function was assessed using an in vitro tube formation assay, demonstrating that immortalized HUVECs retained typical endothelial tube-forming capacity (Figure 5)
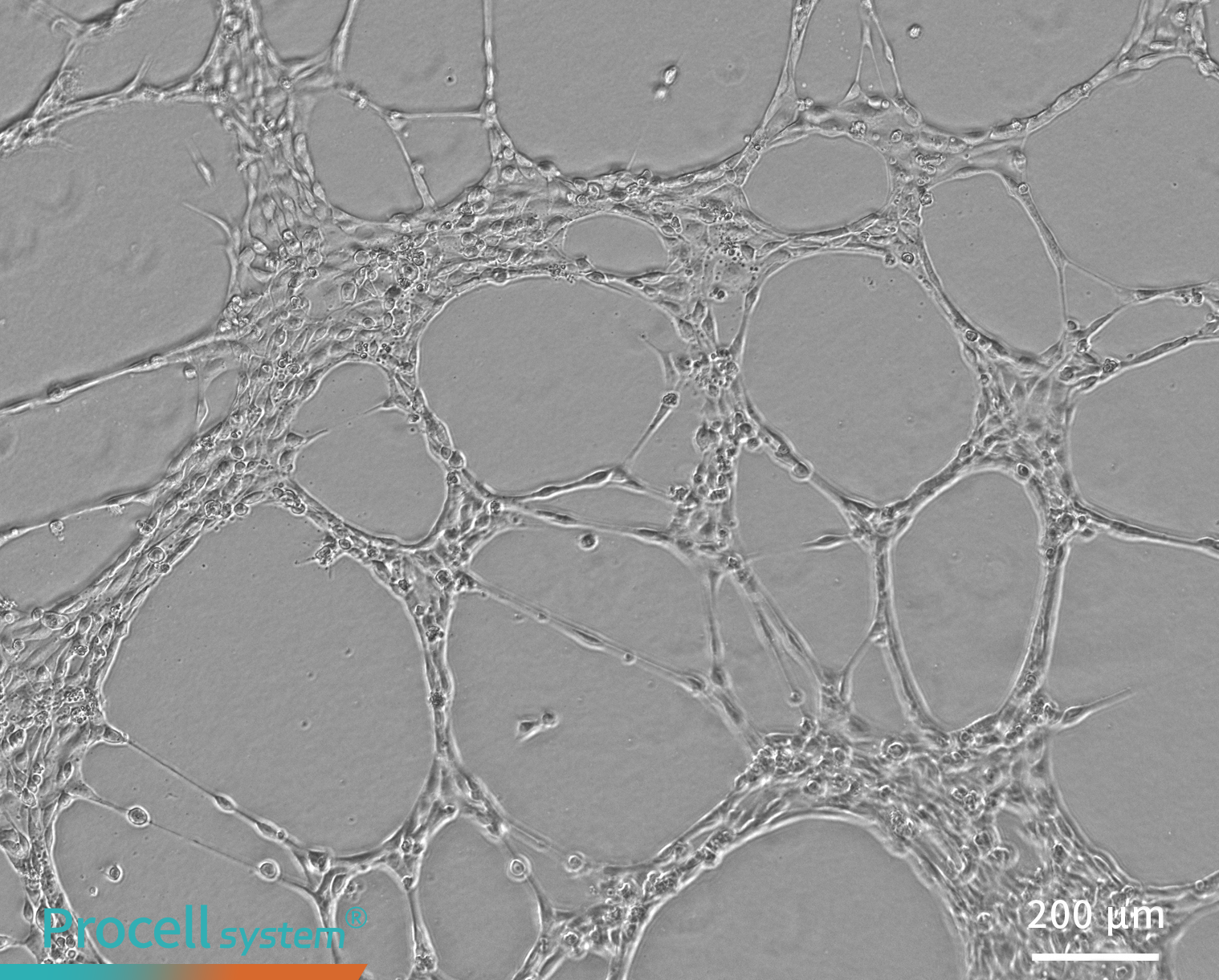
Figure 5. Tube formation assay of immortalized HUVECs in vitro
III. Key Considerations for Primary Cell Immortalization
1. Polybrene Toxicity
Polybrene improves transduction efficiency but can be cytotoxic. For sensitive cells, promptly replace the medium after infection.
2. Minimize Freeze-thaw Cycles
Freeze-thaw cycles reduce viral titer. Aliquot stocks as needed and avoid repeated thawing. Re-titer viruses stored for longer than 6 months before use.
3. Enhancing Transduction in Refractory Cells
For hard-to-transduce cells (e.g., dendritic cells, DCs), use sequential infections: after 24 h, replace the medium with fresh virus for a second round of infection.
4. Functional Validation
Immortalized cells provide unlimited proliferation and stability but may not fully recapitulate primary cell behavior. Confirm their phenotype and function before use.
5. Passage Control
Extended passaging can cause genetic instability and mutation accumulation, altering phenotype and function. Monitor cell status regularly to ensure data reliability.
Prev: A Practical Guide to Antibiotics in Cell Culture
Next: Primary Cell Immortalization: Protocols for Culture, Cryopreservation, and Resuscitation


